|
Centre of Inflammation and Metabolism
CIM |
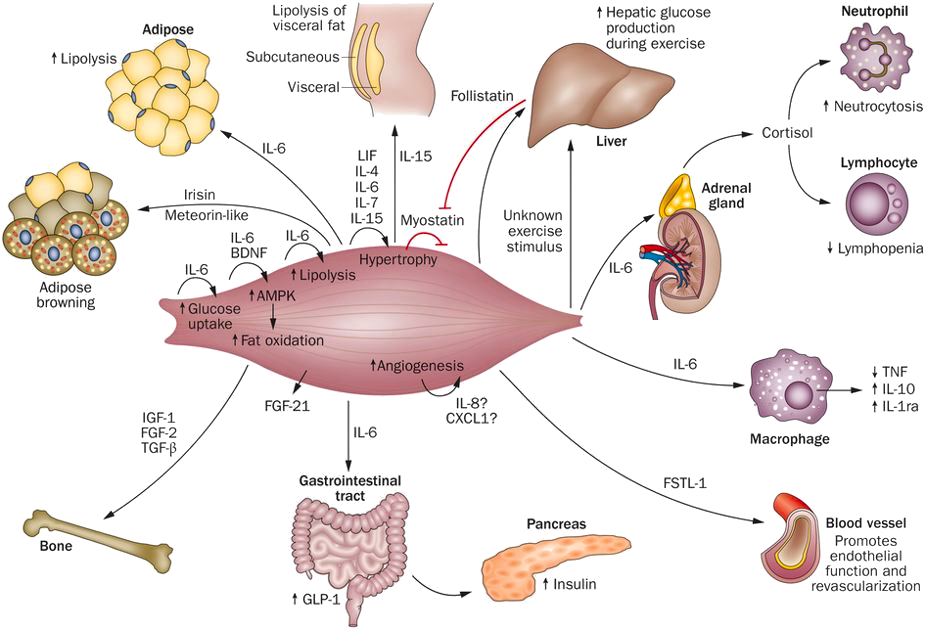
The Centre of Inflammation and Metabolism (CIM) was based on a 5+5-year grant (2005-2015) from the Danish National Research Foundation (DNRF). CIM was founded on the high-risk, innovative idea that skeletal muscle is a secretory organ with the capacity to produce so called “myokines” i.e. cytokines and other peptides that are produced, expressed, and released by muscle fibers and exert either autocrine, paracrine or endocrine effects. We further hypothesized that such myokines would play important roles in cell-to-cell and organ-to-organ cross talk. During the CIM period, we contributed to establish muscle as a secretory organ with the capacity to produce hundreds of myokines. The myokine concept provides a conceptual basis and a new paradigm for understanding how muscles communicate both with other organs and within the muscle itself.
Skeletal muscle produces and releases hundreds of myokines that play a role in muscle-organ cross talk and form a conceptual basis for our understanding of how exercise training may impact diseases with many different phenotypical presentations. The 10 years as a DNRF Center of Excellence made it possible, by combining the efforts of a large group of talented and dedicated researchers and students, to create a unique research concept, linking a molecular approach with whole body in vivo metabolic functions in intact human beings. CIM represented a true multi-disciplinary research environment with an unconventional strategy that was classified “from-man-to-molecule”. We performed integrative physiological research in humans, using invasive methodologies, but with the aim of unravelling mechanisms of action at a molecular level. The fact that the research performed in CIM has had major health aspects related to protection from and treatment of chronic diseases and life style-related disorders, further stimulated us to establish in 2013 the Centre for Physical Activity Research (CFAS) as a true translational research unit focusing on exercise as potent medicine for people with non-communicable diseases. CFAS is based on a center grant from TrygFonden. To learn more, please, visit CFAS at aktivsundhed.dk |
|
| ©
2024 Centre of Inflammation and Metabolism www.inflammation-metabolism.dk |
|